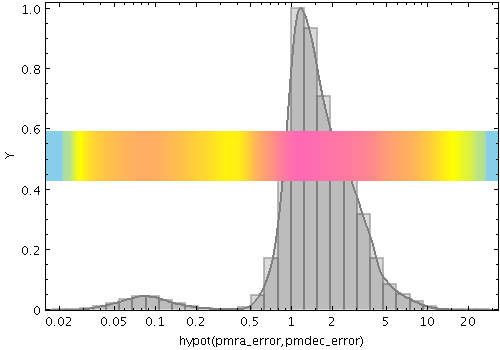
densogram
Represents smoothed density of data values along the horizontal axis using a colourmap. This is like a Kernel Density Estimate (smoothed histogram with bins 1 pixel wide), but instead of representing the data extent vertically as bars or a line, values are represented by a fixed-size pixel-width column of a colour from a colour map. A smoothing kernel, whose width and shape may be varied, is applied to each data point.
A weighting may be applied to the calculated levels
by supplying the
weight
coordinate.
In this case you can choose how these weights are aggregated
in each pixel bin using the
combine
parameter.
The result is something like a smoothed version of the
corresponding weighted histogram.
Note that some combinations of the available parameters
(e.g. a normalised cumulative median-aggregated KDE)
may not make much visual sense.
This is a rather unconventional way to represent density data, and this plotting mode is probably not very useful. But hey, nobody's forcing you to use it.
Usage Overview:
layerN=densogram colorN=<rrggbb>|red|blue|... smoothN=+<width>|-<count>
sidewaysN=true|false
kernelN=square|linear|epanechnikov|cos|cos2|gauss3|gauss6
densemapN=<map-name>|<color>-<color>[-<color>...]
denseclipN=<lo>,<hi> denseflipN=true|false
densequantN=<number> densesubN=<lo>,<hi>
densefuncN=log|linear|histogram|...
cumulativeN=none|forward|reverse sizeN=<pixels>
posN=<fraction> xN=<num-expr> weightN=<num-expr>
inN=<table> ifmtN=<in-format> istreamN=true|false
icmdN=<cmds>
All the parameters listed here
affect only the relevant layer,
identified by the suffix
N.
Example:

stilts plot2plane in=tgas_source.fits x=hypot(pmra_error,pmdec_error)
xscale=log normalise=maximum
color=grey layer1=histogram layer2=kde
layer3=densogram densemap3=skyblue-yellow-hotpink densefunc3=log
size3=50 pos3=0.5
colorN = <rrggbb>|red|blue|... (Color)
The standard plotting colour names are
red, blue, green, grey, magenta, cyan, orange, pink, yellow, black, light_grey, white.
However, many other common colour names (too many to list here)
are also understood.
The list currently contains those colour names understood
by most web browsers,
from AliceBlue to YellowGreen,
listed e.g. in the
Extended color keywords section of
the CSS3 standard.
Alternatively, a six-digit hexadecimal number RRGGBB
may be supplied,
optionally prefixed by "#" or "0x",
giving red, green and blue intensities,
e.g. "ff00ff", "#ff00ff"
or "0xff00ff" for magenta.
[Default: red]
cumulativeN = none|forward|reverse (Cumulation)
forward/reverse
the histogram bars plotted are calculated
cumulatively;
each bin includes the counts from all previous bins
working up/down the independent axis.
Note that setting cumulative plotting may not make much sense with some other parameter values, for instance averaging aggregation modes.
For reasons of backward compatibility,
the values true and false
may be used as aliases for
forward and none.
The available options are:
none: The value plotted for each bin uses the samples accumulated in that bin.forward: The value plotted for each bin uses the samples accumulated all the way from negative infinity to that bin.reverse: The value plotted for each bin uses the samples accumulated all the way from positive infinity to that bin.[Default: none]
denseclipN = <lo>,<hi> (Subrange)
If the full range 0,1 is used,
the whole range of colours specified by the selected
shader will be used.
But if for instance a value of 0,0.5 is given,
only those colours at the left hand end of the ramp
will be seen.
If the null (default) value is chosen,
a default clip will be used.
This generally covers most or all of the range 0-1
but for colour maps which fade to white,
a small proportion of the lower end may be excluded,
to ensure that all the colours are visually distinguishable
from a white background.
This default is usually a good idea if the colour map
is being used with something like a scatter plot,
where markers are plotted against a white background.
However, for something like a density map when the whole
plotting area is tiled with colours from the map,
it may be better to supply the whole range
0,1 explicitly.
denseflipN = true|false (Boolean)
[Default: false]
densefuncN = log|linear|histogram|... (Scaling)
The available options are:
log: Logarithmic scaling, positive values onlylinear: Linear scalinghistogram: Scaling follows data distribution, with linear axishistolog: Scaling follows data distribution, with logarithmic axis, positive values onlyasinh: Asinh scaling, wide dynamic range for both positive and negative valuessqrt: Square root scalingsquare: Square scalingacos: Inverse cosine Scalingcos: Cosine ScalingFor all these options,
the full range of valid data values is used,
and displayed on the colour bar
if applicable (though it can be restricted using the densesub option)
Note that logarithmic-based options
Log and
HistoLog
will ignore non-positive values.
The Linear,
Log,
Square and
Sqrt options
just apply the named function to the full data range.
The histogram options on the other hand use a scaling function
that corresponds to the actual distribution of the data, so that
there are about the same number of points (or pixels, or whatever
is being scaled) of each colour.
The histogram options are somewhat more expensive,
but can be a good choice if you are exploring data whose
distribution is unknown or not well-behaved over
its min-max range.
The Histogram and
HistoLog options both
assign the colours in the same way, but they display the colour
ramp with linear or logarithmic annotation respectively;
the HistoLog option also ignores
non-positive values.
[Default: linear]
densemapN = <map-name>|<color>-<color>[-<color>...] (Shader)
A mixed bag of colour ramps are available
as listed in Section 8.7:
inferno,
magma,
plasma,
viridis,
cividis,
cubehelix,
sron,
rainbow,
rainbow2,
rainbow3,
pastel,
cosmic,
ember,
gothic,
rainforest,
voltage,
bubblegum,
gem,
chroma,
sunset,
neon,
tropical,
accent,
gnuplot,
gnuplot2,
specxby,
set1,
paired,
hotcold,
guppy,
iceburn,
redshift,
pride,
rdbu,
piyg,
brbg,
cyan-magenta,
red-blue,
brg,
heat,
cold,
light,
greyscale,
colour,
standard,
bugn,
bupu,
orrd,
pubu,
purd,
painbow,
huecl,
infinity,
hue,
intensity,
rgb_red,
rgb_green,
rgb_blue,
hsv_h,
hsv_s,
hsv_v,
yuv_y,
yuv_u,
yuv_v,
scale_hsv_s,
scale_hsv_v,
scale_yuv_y,
mask,
blacker,
whiter,
transparency.
Note:
many of these, including rainbow-like ones,
are frowned upon by the visualisation community.
You can also construct your own custom colour map
by giving a sequence of colour names separated by
minus sign ("-") characters.
In this case the ramp is a linear interpolation
between each pair of colours named,
using the same syntax as when specifying
a colour value.
So for instance
"yellow-hotpink-#0000ff"
would shade from yellow via hot pink to blue.
[Default: inferno]
densequantN = <number> (Double)
If left blank, the colour map is nominally continuous (though in practice it may be quantised to a medium-sized number like 256).
densesubN = <lo>,<hi> (Subrange)
The default value "0,1" therefore has
no effect.
The range could be restricted to its lower half
with the value 0,0.5.
[Default: 0,1]
icmdN = <cmds> (ProcessingStep[])
inN.
The value of this parameter is one or more of the filter
commands described in Section 6.1.
If more than one is given, they must be separated by
semicolon characters (";").
This parameter can be repeated multiple times on the same
command line to build up a list of processing steps.
The sequence of commands given in this way
defines the processing pipeline which is performed on the table.
Commands may alternatively be supplied in an external file,
by using the indirection character '@'.
Thus a value of "@filename"
causes the file filename to be read for a list
of filter commands to execute. The commands in the file
may be separated by newline characters and/or semicolons,
and lines which are blank or which start with a
'#' character are ignored.
A backslash character '\' at the end of a line
joins it with the following line.
ifmtN = <in-format> (String)
inN.
The known formats are listed in Section 5.1.1.
This flag can be used if you know what format your
table is in.
If it has the special value
(auto) (the default),
then an attempt will be
made to detect the format of the table automatically.
This cannot always be done correctly however, in which case
the program will exit with an error explaining which
formats were attempted.
This parameter is ignored for scheme-specified tables.
[Default: (auto)]
inN = <table> (StarTable)
-",
meaning standard input.
In this case the input format must be given explicitly
using the ifmtN
parameter.
Note that not all formats can be streamed in this way.:<scheme-name>:<scheme-args>.<" character at the start,
or a "|" character at the end
("<syscmd" or
"syscmd|").
This executes the given pipeline and reads from its
standard output.
This will probably only work on unix-like systems.istreamN = true|false (Boolean)
inN parameter
will be read as a stream.
It is necessary to give the
ifmtN parameter
in this case.
Depending on the required operations and processing mode,
this may cause the read to fail (sometimes it is necessary
to read the table more than once).
It is not normally necessary to set this flag;
in most cases the data will be streamed automatically
if that is the best thing to do.
However it can sometimes result in less resource usage when
processing large files in certain formats (such as VOTable).
This parameter is ignored for scheme-specified tables.
[Default: false]
kernelN = square|linear|epanechnikov|cos|cos2|gauss3|gauss6 (Kernel1dShape)
The available options are:
square: Uniform value: f(x)=1, |x|=0..1linear: Triangle: f(x)=1-|x|, |x|=0..1epanechnikov: Parabola: f(x)=1-x*x, |x|=0..1cos: Cosine: f(x)=cos(x*pi/2), |x|=0..1cos2: Cosine squared: f(x)=cos^2(x*pi/2), |x|=0..1gauss3: Gaussian truncated at 3.0 sigma: f(x)=exp(-x*x/2), |x|=0..3gauss6: Gaussian truncated at 6.0 sigma: f(x)=exp(-x*x/2), |x|=0..6[Default: epanechnikov]
posN = <fraction> (Double)
[Default: 0.05]
sidewaysN = true|false (Boolean)
false,
the quantity being accumulated is on the the horizontal axis
and the frequency is represented vertically as usual.
If set true the quantity accumulated
is on the vertical axis,
and the frequency is represented horizontally,
so that the chart is displayed reflected in the X=Y line.
[Default: false]
sizeN = <pixels> (Integer)
[Default: 12]
smoothN = +<width>|-<count> (BinSizer)
If the supplied value is a positive number it is interpreted as a fixed width in the data coordinates of the X axis (if the X axis is logarithmic, the value is a fixed factor). If it is a negative number, then it will be interpreted as the approximate number of smooothing widths that fit in the width of the visible plot (i.e. plot width / smoothing width). If the value is zero, no smoothing is applied.
When setting this value graphically, you can use either the slider to adjust the bin count or the numeric entry field to fix the bin width.
[Default: -100]
weightN = <num-expr> (String)
The value is a numeric algebraic expression based on column names as described in Section 10.
xN = <num-expr> (String)
The value is a numeric algebraic expression based on column names as described in Section 10.